mdCTP, 5-mdCTP
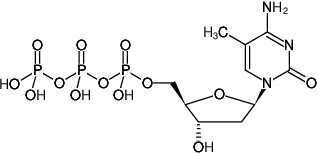
5-Methyl-2'-deoxycytidine-5'-triphosphate, Sodium salt
| Cat. No. | Amount | Price (EUR) | Buy / Note |
|---|---|---|---|
| NU-1125S | 10 μl (100 mM) | 79,80 | Add to Basket/Quote Add to Notepad |
| NU-1125L | 5 x 10 μl (100 mM) | 332,50 | Add to Basket/Quote Add to Notepad |
For general laboratory use.
Shipping: shipped on gel packs
Storage Conditions: store at -20 °C
Short term exposure (up to 1 week cumulative) to ambient temperature possible.
Shelf Life: 12 months after date of delivery
Molecular Formula: C10H18N3O13P3 (free acid)
Molecular Weight: 481.18 g/mol (free acid)
Exact Mass: 481.01 g/mol (free acid)
CAS#: 22003-12-9
Purity: ≥ 95 % (HPLC)
Form: solution in water
Color: colorless to slightly yellow
Concentration: 100 mM - 110 mM
pH: 7.5 ±0.5
Spectroscopic Properties: λmax 277 nm, ε 9.0 L mmol-1 cm-1 (Tris-HCl pH 7.5)
Applications:
Incorporation into DNA by
- PCR with Taq polymerase in-house data, [1-2]
Description:
5-methylated DNA probes can be used as methylation reference fragment [1-2] or for pull-down of 5-hmC binding proteins from cellular lysate[3].
BIOZ Product Citations:
Selected References:
[1] Szwagierczak et al. (2011) Characterization of PvuRts1I endonuclease as a tool to investigate genomic 5-hydroxymethylcytosine. Nucl. Acids Res. 39(12):5149.
[2] Szwagierczak et al. (2010) Sensitive enzymatic quantification of 5-hydroxymethylcytosine in genomic DNA. Nucl. Acids Res. 38(19):e181.
[3] Lafaye et al. (2014) DNA binding of the p21 repressor ZBTB2 is inhibited by cytosine hydroxymethylation. Biochem. Biophys. Res. Commun. 446:341.
Kaito et al. (2001) Activation of the maternally preset program of apoptosis by microinjection of 5-aza-2'-deoxycytidine and 5-methyl-2'-deoxycytidine-5'-triphosphate in Xenopus laevis embryos. Dev Growth Differ. 43 (4):383.
Wu et al. (2007) Crystal structure of RS21-C6, involved in nucleoside triphosphate pyrophosphohydrolysis. J. Mol. Biol. 367:1405.
Dezhurov et al. (2005) A new highly efficient photoreactive analogue of dCTP. Synthesis, characterization, and application in photoaffinity modification of DNA binding proteins. Bioconjug. Chem. 16 (1):215.
Song et al. (2015) Human dCTP pyrophosphatase 1 promotes breast cancer cell growth and stemness through the modulation on 5-methyl-dCTP metabolism and global hypomethylation. Oncogenesis 4: e159.
Frauer et al. (2009) A versatile non-radioactive assay for DNA methyltransferase activity and DNA binding. Nucleic Acids Research 37 (3):e22.
Zeschnigk et al. (2009) Massive parallel bisulfite sequencing of CG-rich DNA fragments reveals that methylation of many X-chromosomal CpG islands in female blood DNA is incomplete. Human Molecular Genetics 18 (8):1439.
Holliday et al. (2002) DNA methylation and epigenetic inheritance. Methods. 27 (2):179.
Asamizu et al. (1999) A large scale structural analysis of cDNAs in a unicellular green alga, Chlamydomonas reinhardtii. I. Generation of 3433 non-redundant expressed sequence tags. DNA Res. 6 (6):369.
Ito et al. (1998) Systematic method to obtain novel genes that are regulated by mi transcription factor: impaired expression of granzyme B and tryptophan hydroxylase in mi/mi cultured mast cells. Blood 91 (9):3210.
Lefaucheur et al. (1998) Evidence for three adult fast myosin heavy chain isoforms in type II skeletal muscle fibers in pigs. J. Anim. Sci. 76 (6):1584.
Wong et al. (1997) A novel method for producing partial restriction digestion of DNA fragments by PCR with 5-methyl-CTP. Nucleic Acids Res. 25 (20):4169.
Chen et al. (1993) Direct induction of DNA hypermethylation in sea urchin embryos by microinjection of 5-methyl dCTP stimulates early histone gene expression and leads to developmental arrest. Dev Biol. 155 (1):75.
[3] Lafaye et al. (2014) DNA binding of the p21 repressor ZBTB2 is inhibited by cytosine hydroxymethylation. Biochem. Biophys. Res. Commun. 446:341.
Nelson et al. (1993) Restriction endonuclease cleavage of 5-methyl-deoxycytosine hemimethylated DNA at high enzyme-to-substrate ratios. Nucl. Acid. Res. 21 (3):681.