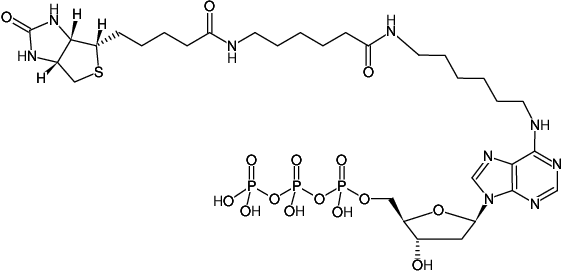
Biotin-14-N6-(6-Aminohexyl)-dATP, Triethylammonium salt
Bio-14-dATP
| Cat. No. | Amount | Price (EUR) | Buy / Note |
|---|---|---|---|
| NU-835-BIO14-S | 200 μl (1 mM) | 122,80 | Add to Basket/Quote Add to Notepad |
| NU-835-BIO14-L | 5 x 200 μl (1 mM) | 375,70 | Add to Basket/Quote Add to Notepad |
For general laboratory use.
Shipping: shipped on gel packs
Storage Conditions: store at -20 °C
Short term exposure (up to 1 week cumulative) to ambient temperature possible.
Shelf Life: 12 months after date of delivery
Molecular Formula: C32H54N9O15P3S (free acid)
Molecular Weight: 929.81 g/mol (free acid)
Exact Mass: 929.27 g/mol (free acid)
Purity: ≥ 95 % (HPLC)
Form: filtered solution (30 kDa) in 10 mM Tris-HCl
Color: colorless to slightly yellow
Concentration: 1.0 mM - 1.1 mM
pH: 7.5 ±0.5
Spectroscopic Properties: λmax 266 nm, ε 16.2 L mmol-1 cm-1 (Tris-HCl pH 7.5)
Applications:
Incorporation into DNA/cDNA by
- Nick Translation with DNAse I/ DNA Polymerase I [1] & in-house data
- Primer Extension with Klenow fragment [2]
Description:
Biotin-14-dATP is enzymatically incorporated into DNA/cDNA as substitute for its natural counterpart dATP. The resulting Biotin-labeled DNA/cDNA probes are subsequently detected using streptavidin conjugated with horseradish peroxidase (HRP), alkaline phosphatase (AP), a fluorescent dye or agarose/magnetic beads. Optimal substrate properties for Nick Translation are ensured by a 14-atom linker attached to the N6 position of adenine. For PCR incorporation experiments e.g. with Taq polymerase Biotin-11-dATP (#NU-1175-BIOX) is recommended whose Biotin moiety is attached to the N7-Deaza position of adenine via a 11-atom linker.
Recommended Biotin-14-dATP/dATP ratio for Nick Translation: 50% Biotin-14-dATP/ 50% dATP
Please note: The optimal final concentration of Biotin-14-dATP may very depending on the application and assay conditions. For optimal product yields and high incorporation rates an individual optimization of the Biotin-14-dATP/dATP ratio is recommended.
Related products:
BIOZ Product Citations:
Selected References:
[1] Gebeyehu et al. (1987) Novel biotinylated nucleotide-analogs for labeling and colorimetric detection of DNA. Nucleic Acids Res. 15 (21):4513.
[2] Nagano et al. (2015) Single-cell Hi-C for genome-wide detection of chromatin interactions that occur simultaneously in a single cell. Nature Protocols 10 (12):1987.
Mumbach et al. (2016) HiChIP: Efficient and sensitive analysis of protein-directed genome architecture Nature Protocols 13 (11):919.