Apoptosis is the process of an intracellular death program leading to characteristic biochemical and morphological changes within a cell that consequently result in cell death[1]. A failure of cells to undergo apoptosis is a common feature of many cancers [2] thus investigation of apoptosis inducing mechanisms is of particular importance for cancer research and requires suitable detection methods.
A hallmark of late apoptosis is extensive genomic DNA fragmentation that generates a multitude of DNA double-strand breaks (DSBs) with accessible 3'-hydroxyl (3'-OH) groups. This characteristic forms the basis for a well-established apoptosis detection method: Terminal deoxynucleotidyl transferase dUTP Nick End Labeling (TUNEL) assay[3].
TUNEL assays identify apoptotic cells by the terminal deoxynucleotidyl transferase (TdT)-mediated addition of labeled (X) deoxyuridine triphosphate nucleotides (X-dUTPs) to the 3’-OH end of DNA strand breaks that are subsequently visualized depending on the introduced label (Fig. 1), thus serving as parameter for the percentage of apoptotic cells within the analyzed cell population.
The assay sensitivity strongly depends on the incorporation efficiency of the modified dUTP that is influenced by size/bulkiness of the attached label. A number of fluorescently labeled dUTPs → Probes & Epigenetics, biotinylated dUTPs → Probes & Epigenetics or digoxigenylated dUTPs → Probes & Epigenetics are generally suitable substrates for TdT, but dUTP labeled with smaller labels such as bromine (BrdUTP) or an alkyne group (EdUTP) have been demonstrated to exhibit a higher incorporation efficiency and thus higher sensitivity in TUNEL assays probably due to minimal sterical hindrance[3,4,5].
While incorporation of BrdUTP is detected by specific fluorophore or reporter enzyme labeled antibodies (Fig. 1A), detection of EdUTP is achieved by covalent conjugation of an azide-containing fluorophore → Click Chemistry via Cu(I) catalyzed alkyne-azide click chemistry (CuAAC) (Fig. 1B).
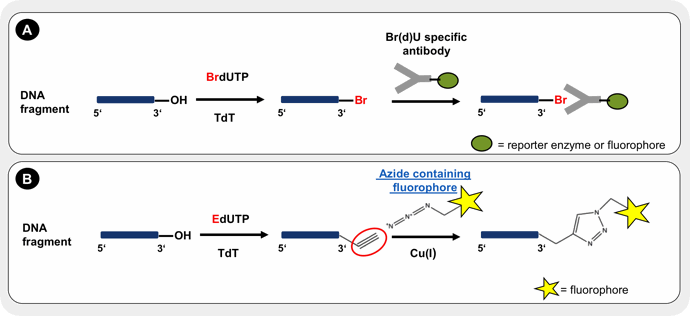
Figure 1: The principle of TUNEL assay relies on terminal deoxynucleotidyl transferase (TdT)-mediated addition of a modified dUTP (X-dUTP) to 3’-OH ends of DNA fragments that are generated as a result of apoptosis induction. To avoid the loss of fragmented DNA and to allow enzyme and nucleotide entrance, cells need to be fixed and subsequently permeabilized prior to the labeling reaction. Incorporated bromoylated dUTP (BrdUTP) is detected by specific antibody conjugates with a reporter enzyme or fluorescent dye (A). The incorporation of alkyne-containing dUTP (EdUTP) is visualized by Cu(I)-catalyzed alkyne-azide click chemistry (CuAAC) with an azide containing fluorphore → Click Chemistry (B).
[1] Kerr et al. (1972) Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br. J. Cancer 26 (4):239.
[2] Lowe et al. (2000) Apoptosis in Cancer. Carcinogenesis 21 (3):485.
[3] Darzynkiewicz et al. (2008) Analysis of apoptosis by cytometry using TUNEL assay. Methods 44 (3):250.
[4] Li et al. (1995) Labelling DNA strand breaks with BrdUTP: Detection of apoptosis and cell proliferation. Cell proliferation 28 (11):571.
[5] Li et al. (1995) Single–step procedure for labeling DNA strand breaks with fluorescein- or BODIPY-conjugated deoxynucleotides: Detection of apoptosis and bromdeoxyuridine incorporation. Cytometry 20:172.