The dynamics of global protein synthesis (both spatial and temporal) is an essential parameter to characterize the cellular response under various physiological and pathological conditions. Protein synthesis has traditionally been monitored by metabolic labeling with 35S-methionine that competes with natural methionine for the incorporation into newly synthesized proteins. The detection is limited to autoradiography.
The cell-permeable methionine-analogs 4-Azido-L-homoalanine (L-AHA) and L-Homopropargylglycine (HPG) provide a non-radioactive alternative to analyze the global protein synthesis in cell culture. They are randomly incorporated instead of methionine during translation leading to full-length, azide-labeled or alkyne-modified proteins, respectively[1,2,3](Fig. 1).
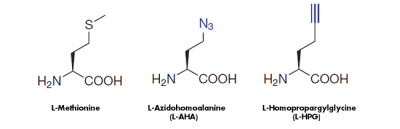
Figure 1: Methionine analogs are suitable to incorporate azide groups a href="/click-chemistry/click-reagents-by-chemistry/azide-reagents/amino-acids/clk-aa005-4-azido-l-homoalanine-hcl-l-aha">(L-AHA) or alkyne groups (L-HPG) into nascent proteins. Visualization of the incorporated azide groups (L-AHA-based labeling) is performed via Cu(I)-catalyzed Click Chemistry with Alkynes of fluorescent dyes or via Cu-free Click Chemistry with DBCO-containing fluorescent dyes or DBCO-containing Biotin. Visualiation of Alkyne groups (HPG-based labeling) can be performed with Cu(I)-catalayzed Click Chemistry only.
[1] Dieck et al. (2012) Metabolic Labeling with Noncanonical Amino Acids and Visualisation by Chemoselective Fluorescent Tagging. Current Protocols in Cell Biology 7:7.11.1.
[2] Kiick et al. (2002) Incorporation of azides into recombinant proteins for chemoselective modification by the Staudinger ligation. Proc. Natl. Acad. Sci. USA 99 (1):19.
[3] Dieterich et al. (2010) In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons. Nature Neuroscience 13 (7):897.