NTC or clonNAT
sterile ready-to-go stock solution
| Cat. No. | Amount | EUR / ml | Price (EUR) | Buy / Note |
|---|---|---|---|---|
| AB-101S | 1 ml (100 mg/ml) | 45,00 | 45,00 | Add to Basket/Quote Add to Notepad |
| AB-101L | 5 ml (100 mg/ml) | 34,40 | 172,00 | Add to Basket/Quote Add to Notepad |
| AB-101-10ML | 10 ml (100 mg/ml) | 31,50 | 315,00 | Add to Basket/Quote Add to Notepad |
| AB-101-50ML | 50 ml (100 mg/ml) | 23,10 | 1.155,00 | Add to Basket/Quote Add to Notepad |
For general laboratory use. Not intended for human or animal diagnostic or therapeutic uses.
Shipping: shipped on gel packs
Storage Conditions: Store at -20 °C
Shelf Life: 12 months
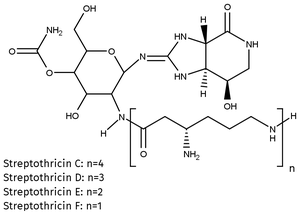
Molecular Formula: C19H34N8O8 ∙ H2SO4 (Streptothricin F)
Molecular Weight: 600.6 g/mol (Streptothricin F)
CAS#: 96736-11-7
Form: liquid
Color: beige
Concentration: 100 mg/ml of sterile filtrated Nourseothricin in water
pH: 6.3
Description:
Nourseothricin is a mixturte of Streptothricins C, D, E and F and can be used as selection antibiotic for a broad spectrum of pro- and eukaryotic organisms (i.e. Gram-positive and Gram-negative bacteria, yeast, filamentous fungi, protozoa, microalgae, plants and many more).
Selection of recombinant strains is based on inactivation of Nourseothricin by mono-acetylation of the ß-amino group of the ß-lysine residue by Nourseothricin N-acetyltransferase, the product of the sat1 or nat1 genes.
Selection:
For selection of recombinant Leishmania strains Nourseothricin is added to the growth medium to a final concentration of 100 μg/ml.
For selection of other species please refer to the product page.
BIOZ Product Citations:
Selected References:
[1] Goldstein et al. (1999) Three New Dominant Drug Resistance Cassettes for Gene Disruption in Saccharomyces cerevisiae. Yeast 15: 1541
[2] Kojic et al. (2000) Shuttle vectors for genetic manipulations in Ustilago maydis. Can. J. Microbiology 46: 333
[3] Werner et al. (2001) Aminoglycoside-Streptothricin Resistance Gene Cluster aadE–sat4–aphA-3 Disseminated among multiresistant Isolates of Enterococcus faecium. Antimicrob. Agents Chemotherapy 45: 3267
[4] Hoff et al. (2009) Homologous recombination in the antibiotic producer Penicillium chrysogenum: strain ΔPcku70 shows up-regulation of genes from the HOG pathway. Appl. Microbiol. Biotechnol. 85:1081
[5] Kochupurakkal & Iglehart (2013) Nourseothricin N-Acetyl Transferase: A Positive Selection Marker for Mammalian Cells. PLoS One 8: e68509
[6] Ramos et al. (2013) Functional genomics tools to decipher the pathogenicity mechanisms of the necrotrophic fungus Plectosphaerella cucumerina in Arabidopsis thaliana. Molecular Plant Pathology 14: 44
[7] Schubert et al. (2013) Agrobacterium-mediated transformation of the white-rot fungus Physisporinus vitreus. J. Microbiol. Meth. 95: 251
[8] Buhmann et al. (2014) A Tyrosine-Rich Cell Surface Protein in the Diatom Amphora coffeaeformis Identified through Transcriptome Analysis and Genetic Transformation. PLOS one 9: e110369
[9] Kraeva et al. (2015) Leptomonas seymouri: Adaptations to the Dixenous Life Cycle Analyzed by Sequencing, Transcriptome Profiling and Co-infection with Leishmania donovani. PLOS Pathogens 11: e1005127
[10] Paschke et al. (2018) Rapid and efficient genetic engineering of both wild type and axenic strains of Dictyostelium discoideum. PLoS One 13: e0196809