To study:
Crucial cellular processes such as transport, motion and division are orchestrated by a highly diverse set of regulatory phosphatases. To trap these enzymes in their various biologically relevant states, non-hydrolyzable analogs of nucleotides are ideally suited. Jena Bioscience has more than two decades of experience in the preparation of non-hydrolyzable nucleotides, supplying researchers with >50 catalog products (see Scheme 1) of this compound class. At the forefront of science1-10 it is our mission to facilitate your work on:
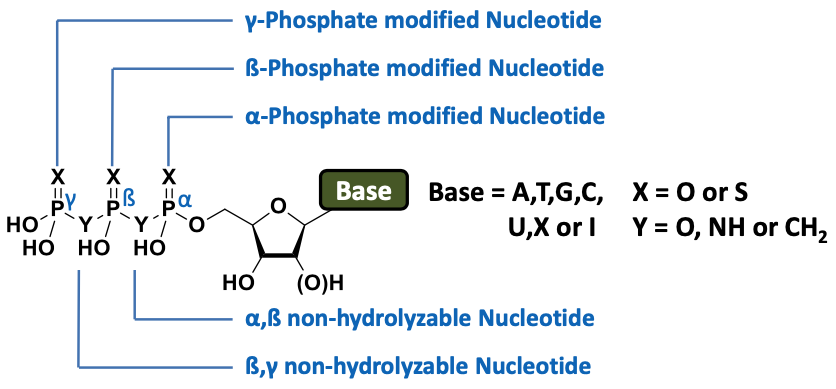
Scheme 1: Overview of Jena Bioscience’s non-hydrolyzable nucleotides. Chemical resistance to hydrolysis can be achieved by replacing bridging oxygens for imido (-NH-) or methylene (-CH2-) moieties. Moreover, phosphorothioates show markedly reduced rates of hydrolysis compared to the corresponding phosphate analogues while preserving a high sterical similarity. (Please click the position of modification topic for ordering information.)
| Interactions | |
|
[1] Waschbüsch et al. (2019) Rab32 interacts with SNX6 and affects retromer-dependent Golgi trafficking. PLoS One 14 (1):e0208889. |
|
| Nucleotide | Applications
|
|
[3] Whitecross et al. (2017) Identification of the Binding Sites on Rab5 and p110beta Phosphatidylinositol 3-kinase. Sci. Rep. 7 (1):16194. |
|
| Nucleotide | Application
|
|
[4] Bodnar et al. (2017) Molecular Mechanism of Substrate Processing by the Cdc48 ATPase Complex. Cell 169 (4):722. |
|
| Nucleotide | Application
|
| Structure and Dynamics | |
|
[5] Parker et al. (2018) K-Ras Populates Conformational States Differently from Its Isoform H-Ras and Oncogenic Mutant K-RasG12D. Structure 26 (6):810. |
|
| Nucleotide | Applications
|
|
[6] Talavera et al. (2018) Phosphorylation decelerates conformational dynamics in bacterial translation elongation factors. Sci. Adv. 4 (3):eaap9714. |
|
| Nucleotide | Application
|
| Kinetics | |
|
[7] Biancucci et al. (2018) The bacterial Ras/Rap1 site-specific endopeptidase RRSP cleaves Ras through an atypical mechanism to disrupt Ras-ERK signaling. Sci. Signal. 11 (550). pii: eaat8335. |
|
| Nucleotide | Application
|
|
[9] O'Donnell et al. (2017) Timing and Reset Mechanism of GTP Hydrolysis-Driven Conformational Changes of Atlastin. Structure 25 (7):997. |
|
| Nucleotide | Application
|
|
[10] Yu et al. (2018) Cryo-EM Structures of MDA5-dsRNA Filaments at Different Stages of ATP Hydrolysis. Mol. Cell. 72 (6):999. |
|
| Nucleotide | Application
|

You haven't found the particular nucleotiode analog you're looking for? You have questions or need further information? Do not hesitate to contact us at nucleotides@jenabioscience.com !