Fluorescent nucleotide conjugates are analytical tools with exceptional versatility. By covering a plethora of fluorescence-based techniques they are applied as drug screening probes, in cytoskeletal imaging and in nucleotide-protein interaction studies. Learn more below!
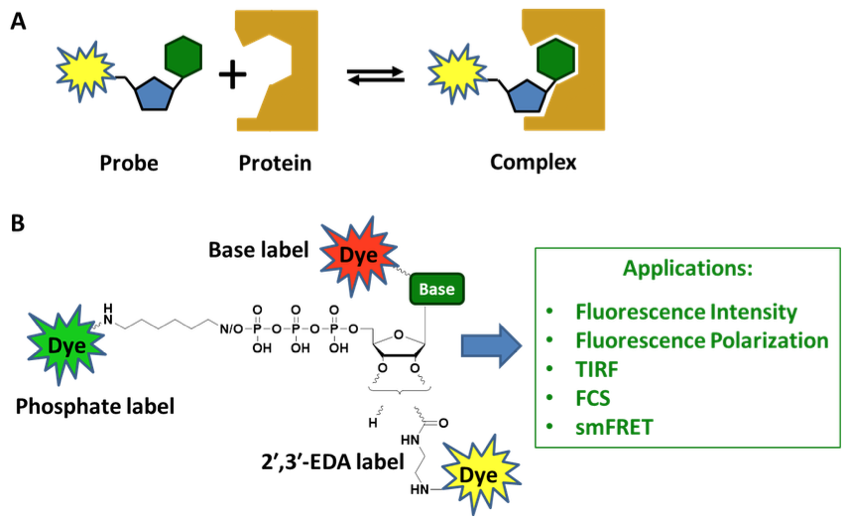
Scheme 1:
A) Binding equilibrium between a fluorescent nucleotide probe and its target protein.
B) A fluorescent dye can be placed at virtually any chemical constituent of a nucleotide, namely phosphate moieties, sugar and base. The corresponding dye conjugates are applied in numerous fluorescence-based assays such as intensity/polarization measurements, total internal reflection fluorescence (TIRF) microscopy, fluorescence correlation spectroscopy (FCS) and single molecule fluorescence resonance energy transfer (smFRET).
Binding interactions between nucleotides and proteins (Scheme 1A) are crucial for stabilizing nucleic acid structures and for regulating ATPases and GTPases, the ubiquitous motors and switches of the cell.
To characterize the biophysical features of nucleotide-protein interactions, highly sensitive fluorescence-based techniques offer a versatile toolbox. Our broad collection of dye-labeled nucleotides covers the entire structural space of suitable labeling positions (Scheme 1B) and has been successfully applied for:

Numerous dye labels can be combined with additional useful nucleotide modifications such as non-hydrolyzable imidodiphosphates, methylenodiphosphates or phosphorothioates. Please inquire at nucleotides@jenabioscience.com !
|
[1] Lang et al. (2017) Actin ADP-ribosylation at Threonine148 by Photorhabdus luminescens toxin TccC3 induces aggregation of intracellular F-actin. Cell. Microbiol. 19 (1):e12636. |
|
| Compounds
EDA-GTP - Cy3 (Cat.No. NU-820-CY3) |
Applications
|
|
[3] Singh et al. (2018) Crystallographic and enzymatic insights into the mechanisms of Mg-ADP inhibition in the A1 complex of the A1AO ATP synthase. J. Struct. Biol. 201 (1):26. |
|
| Compounds
EDA-ATP - ATTO-647N (Cat.No. NU-808-647N) |
Applications
|
|
[6] Evans et al. (2015) Investigating Apoptozole as a Chemical Probe for HSP70 Inhibition. PLoS One 10 (10):e0140006. |
|
| Compounds
N6-(6-Aminohexyl)-ATP - 5-FAM (Cat.No. NU-805-5FM) |
Application
|
|
[9] Wortmann et al. (2017) Cooperative Nucleotide Binding in Hsp90 and Its Regulation by Aha1. Biophys. J. 113 (8):1711. |
|
| Compound
γ-[(6-Aminohexyl)imido]-AppNHp - Atto-647N (Custom Product) |
Application
|
|
[10] Yuan et al. (2016) A novel small-molecule compound disrupts influenza A virus PB2 cap-binding and inhibits viral replication. J. Antimicrob. Chemother. 71 (9):2489. |
|
| Compound | Application
|
|
[11] Lee et al. (2017) Mechanism of SOS PR-domain autoinhibition revealed by single-molecule assays on native protein from lysate. Nat. Commun. 8:15061. |
|
| Compounds
EDA-GDP - ATTO-488 (Cat.No. NU-840-488) |
Application
|