Fluorescent labeling of thiol groups
| Cat. No. | Amount | Price (EUR) | Buy / Note |
|---|---|---|---|
| FP-202-655 | 3 reactions | 670,70 | Add to Basket/Quote Add to Notepad |
For general laboratory use.
Shipping: shipped on gel packs
Storage Conditions: store at -20 °C
store dark, see manual
Shelf Life: 12 months
Spectroscopic Properties:
Excitation maximum: λexc = 663 nm
Emission maximum: λem = 684 nm
Extinction coefficient: εmax = 125,000 mol-1 cm-1
Correction factor: CF280 nm = 0.08
Description:
Fluorescence techniques have become a major tool in biological sciences. Fluorescent proteins such as GFP and DsRed fused to the protein of interest (POI) allow expression analysis and in vivo protein localization. Their application however, is limited due to the large molecular weight of those fusion proteins. Furthermore, the associated molecular biology is tedious and time-consuming.
Small fluorophores, covalently attached to the POI, may help to overcome this problem. The most common labeling method for proteins is amine modification. Amine-reactive fluorophores such as NHS-esters readily react with lysines, present at the surface of the POI, resulting in a stable covalent bond. (Please note: Jena Bioscience offers a ready-to-use kit for labeling of lysine residues with high-quality small fluorophores (Cat.# FP-201) for application in immunochemistry and other assays that require extraordinary stability of the conjugate).
Although fluorescent labeling of lysine residues is a reliable method, the alternative labeling of the thiol group of cysteines is often preferred for investigating structure, function and interactions of the POI. Since many enzymes contain a cysteine in their catalytic domain, protein activity and function may be conveniently investigated by targeting cysteines involved in the catalytic process or binding site.[1,2]
In general, thiol labeling is more specific than amine labeling due to the relatively low abundance of cysteines compared to lysines. Since however, appearance of cysteines and lysines varies among different proteins, the amino acid sequence and tertiary structure of the POI must be analyzed prior to deciding on the labeling strategy.
This Jena Bioscience Protein Labeling Kit is designed for labeling the cysteines of a POI with a small fluorophore resulting in a fluorescent protein-fluorophore conjugate. It contains all reagents required for performing 3 separate labeling reactions of 1 mg of POI.
Content:
Atto 655 Maleimide
1 vial containing 1 mg
Dimethylformamide (DMF)
200 μl
Glutathione (GSH)
3 vials containing 1 mg each
ultra-pure water
1 ml
Storage and Stability:
Upon receipt store the dye at -20 °C. The other components may be stored at 4 °C. If stored as recommended, Jena Bioscience guarantees optimal performance of this product for 12 months.
Protocol:
General notes
The optimal protein concentration is 10 mg/ml, other concentrations are also possible, however, it should be at least 2 mg/ml since labeling efficiency suffers from lower protein concentrations. We recommend using about 1 mg protein per labeling reaction. The optimal pH ranges from 7 to 7.5. At this pH the thiol (sulfhydryl) group is deprotonated to a sufficient degree and readily reacts with the dye-maleimide.
General experimental protocol
Concentration of the conjugate
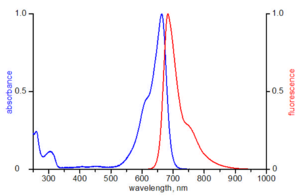
Since there may occur a certain loss of protein during labeling and particularly during purification measuring the concentration of the conjugate is important for further application. The concentration of a protein is commonly determined by measuring its absorbance at 280 nm. As shown in the excitation spectrum of Atto 655 however, the fluorescent dye absorbs at 280 nm as well thereby increasing the ε280 for the conjugate. Therefore, a correction factor (CF) is required to eliminate the contribution of the dye at 280 nm.
The CF of each dye is given in the Spectroscopic Data section, and hence the concentration of the conjugate is calculated according to:
c (mg/ml) = ((A280 - Amax x CF) / ε280 ) x MWprotein
| A280 | absorbance of the conjugate solution measured at 280 nm |
| Amax | absorbance of the conjugate solution measured at λexc |
| λexc, εmax, CF | intrinsic properties of the Atto dye, please refer to Spectroscopic Prop |
| λ280, MW | intrinsic properties of your protein, if not known they can be obtained from web sources such as ExPASy Proteomics Server[3] |
DOL = (Amax x ε280) / ((A280 - Amax x CF) x εmax)
Example for determining the conjugate concentration and the DOL of BSA (Bovine serum albumin), labeled with Atto 488 maleimide:
BSA: ε280 = 42,925 M-1 cm-1, MW = 66,433 Da
Atto 488: λexc = 501 nm, εmax = 90,000 M-1 cm-1, CF = 0.1
After labeling and purification the absorption of the conjugate solution is measured at 280 and 501 nm, respectively.
| - | absorption |
| A280 | 0.29 |
| A501 | 0.5 |
c (mg/ml) = ((A280 - Amax x CF) / ε280 ) x MWprotein
c (mg/ml) = ((0.29 - 0.5 x 0.1) /42,925 ) x 66,433
c (mg/ml) = 0.37
DOL = (Amax x ε280) / ((A280 - Amax x CF) x εmax)
DOL = (0.5 x 42,925) / ((0.29 - 0.5 x 0.1) x 90,000)
DOL = 0.99
In this experiment the concentration of the conjugate is 0.37 mg/ml and most protein molecules are labeled with with approximately one Atto 488 molecule.
Please note that the spectroscopic determination of the concentration and the DOL of the conjugate is not absolutely correct. The spectroscopic characteristics of the free dye are not exactly the same as of the dye bound to the POI. Also, the spectroscopic characteristics of the native protein at 280 nm differ from those of the conjugate.
In general, these changes are negligibly small and therefore, spectroscopic determination of the concentration and the DOL of the conjugate is the most frequently used method[4] .
In addition to the spectroscopic measurements, one may analyze the conjugate by SDS-PAGE and subsequent fluorescence scanning of the gel. Only one band (consisting of the fluorescently labeled protein) should be visible in the fluorescent scan. If there is an additional band at very low molecular weight, the conjugate solution still contains free dye and must be purified again.
Storage of the Conjugate
Protect from light! Store the conjugate just like the unlabeled protein. We recommend dividing the solution into small aliquots and freeze at -20 °C or -80 °C. Avoid repeated freezing and thawing!
Troubleshooting:
Inefficient Labeling
Note that labeling efficiency not only depends on the surrounding conditions but also on the protein characteristics such as tertiary structure and the number of cysteines.
Overlabeling
Reasons for overlabeling can be a high number of cysteines on the surface of your protein.
To prevent overlabeling, decrease the amount of dye or increase the protein concentration. You may also reduce the reaction time.
Purification of the conjugate
The maleimide is unstable in aqueous solutions. Thus, a certain amount of free dye is produced during each labeling reaction and needs to be separated from the conjugate.
If your purified conjugate still contains traces of free dye, apply it to a second purification step. Check the purity of your conjugate by SDS-PAGE.
Increase the amount of GSH proportionally in case you increase the amount of maleimide per labeling reaction. GSH is essential for consuming excess thiol-reactive dye after the labeling reaction is completed. Reactive dye molecules might interfere with the purification of the conjugate and thus, an excess of GSH must be added subsequent to the labeling reaction.
Separation of free dye from the conjugate becomes more difficult at higher concentrations of free dye. The column or membrane might even be blocked by the dye at very high concentrations, so try to optimize the labeling reaction in order to decrease the concentration of free dye. Keep in mind that overall yield suffers from additional purification steps.
BIOZ Product Citations:
Selected References:
[1] Skegro et al. (2007) N-terminal and C-terminal Domains of Arrestin Both Contribute in Binding to Rhodopsin. Photochem. Photobiol. 83:385.
[2] Kamata et al. (2005) Reactive Oxygen Species Promote TNF -Induced Death and Sustained JNK Activation by Inhibiting MAP Kinase Phosphatases. Cell 120:649
[3] www.expasy.ch/tools/protparam.html
[4] Brinkley (1992) A Brief Survey of Methods for Preparing Protein Conjugates with Dyes, Haptens, and Cross-Linking Reagents. Bioconjugate Chem. 3:2.