No login/account required.
Your Basket/Online Quote
Items: 0 (0,00 €)
» Search & Order
» Sign in / Register
Your Basket/Online Quote
Items: 0 (0,00 €)
» Search & Order
» Sign in / Register
Preparation of Cy3-labeled DNA probes by PCR
| Cat. No. | Amount | Price (EUR) | Buy / Note |
|---|---|---|---|
| APP-101-CY3-S | 10 reactions x 20 μl | 145,00 | Add to Basket/Quote Add to Notepad |
| APP-101-CY3-L | 50 reactions x 20 μl | 397,00 | Add to Basket/Quote Add to Notepad |

For general laboratory use.
Shipping: shipped on gel packs
Storage Conditions: store at -20 °C
avoid freeze/thaw cycles, store dark
Shelf Life: 12 months
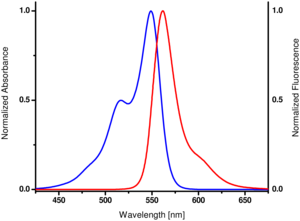
Spectroscopic Properties: λexc 550 nm, λem 570 nm, ε 150.0 L mmol-1 cm-1 (Tris-HCl pH 7.5)
Description:
HighFidelity Cy3 PCR Labeling Kit is designed to produce randomly Cy3-modified DNA probes by PCR. Such probes are ideally suited for Fluorescence in situ hybridization (FISH) and Northern Blot experiments. PCR-based labeling is superior to random-primed labeling with Klenow fragment if template amounts are limited or amplification of a specific DNA fragments is required. Amplification of probes up to 4kbp is feasible.
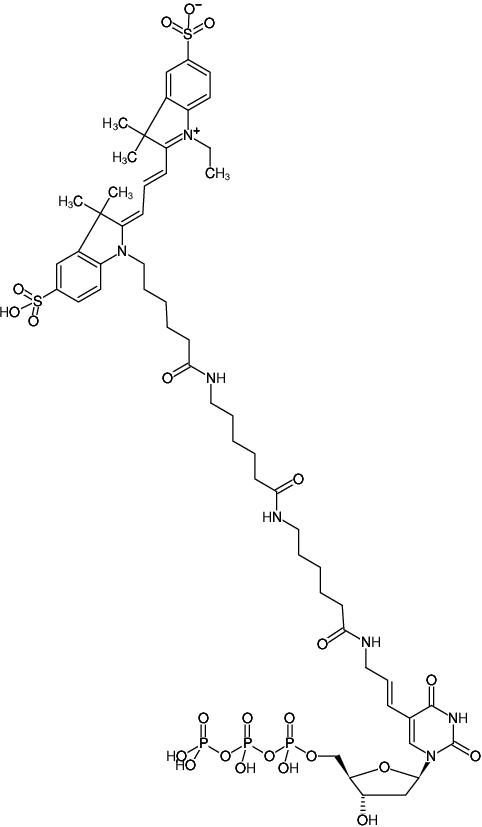
dUTP-XX-Cy3 is efficiently incorporated into DNA as substitute for its natural counterpart dTTP using an optimized reaction buffer and a High Fidelity Polymerase blend consisting of Taq polymerase and a proofreading enzyme. 50 % dUTP-XX-Cy3 substitution typically results in an optimal balance between reaction and labeling efficiency. Individual optimization of dUTP-XX-Cy3/dTTP ratio however, can easily be achieved with the single nucleotide format.
The kit contains sufficient reagents for 10 labeling reactions (S-Pack) or 50 labeling reactions (L-Pack) of 20 μl each (50% dUTP-XX-Cy3 substitution, 100 μM dATP/dGTP/dCTP, 50 μM dTTP, 50 μM dUTP-XX-Cy3).
Content:
High Fidelity Polymerase
in storage buffer with 50% glycerol (v/v)
#APP-101-Cy3-S: 1x 40 μl (100 units, 2.5 units/μl)
#APP-101-Cy3-L: 2x 40 μl (2x 100 units, 2.5 units/μl)
High Fidelity Labeling Buffer
1x 500 μl (10x)
dATP - Solution
1x 20 μl (100 mM)
dGTP - Solution
1x 20 μl (100 mM)
dCTP - Solution
1x 20 μl (100 mM)
dTTP - Solution
1x 20 μl (100 mM)
dUTP-XX-Cy3
#APP-101-Cy3-S: 1x 10 μl (1 mM)
#APP-101-Cy3-L: 5x 10 μl (1 mM)
Lambda DNA
1x 20 μl (100 ng/μl)
500 bp forward primer
1x 20 μl (10 μM)
500 bp reverse primer
1x 20 μl (10 μM)
PCR-grade water
1x 1.2 ml
To be provided by user
DNA template
Primer
DNA purification tools (optional)
1. Preparation of working solutions
1.1 Preparation of 1 mM dATP/dCTP/dGTP working solution
1.2 Preparation of 1 mM dTTP working solution
3. Standard PCR Labeling protocol
The standard protocol is set-up for labeling of a 500 bp DNA fragment. An optimal balance between reaction and labeling efficiency is typically achieved with 50% dUTP-XX-Cy3 substitution following the standard protocol below however, individual optimization might improve results for individual applications.
| Component | Volume | Final concenctration |
| PCR-grade water | X μl | |
| High Fidelity Labeling Buffer (10x) | 2 μl | 1x |
| 1 mM dATP/dCTP/ dGTP working solution (s. 1.1) | 2 μl | 100 μM |
| 1 mM dTTP working solution (s. 1.2) | 1 μl | 50 μM |
| 1 mM dUTP-XX-Cy3 | 1 μl | 50 μM |
| forward primer (10 μM) | X μl | 0.1 - 1 μM (e.g. 0.3 μM 500 bp forward primer) |
| reverse primer (10 μM) | X μl | 0.1 - 1 μM (e.g. 0.3 μM 500 bp reverse primer) |
| template DNA | X μl | 1 - 10 ng genomic DNA (e.g. 1 ng Lambda DNA) |
| High Fidelity Polymerase (2.5 units/μl) | 1 μl | 2.5 units |
| Total volume | 20 μl |
Recommended cycling conditions
| Cycle step | Temperature | Time | Cycles |
| Initial denaturation | 95°C | 2 min | 1x |
| Denaturation Annealing1) Elongation2) | 95°C 58°C 68°C | 20 sec 30 sec 60 sec | 30x |
| Final Elongation | 68°C | 2 min | 1x |
For optimal amplification results and high incorporation rates an individual optimization of the recommended PCR assay and cycling conditions may be necessary for each new primer-template pair.
4. Probe purification:
Probe purification is not required for most hybridization experiments. If a downstream application requires purification (e.g. concentration determination by absorbance measurement) we recommend silica-membrane or gel filtration-based purification.
Related products: Aminoallyl-dUTP-XX-Cy3, #NU-803-XX-Cy3
BIOZ Product Citations: