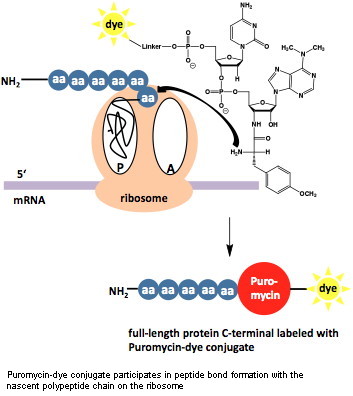
The antibiotic puromycin is a structural analog of aminoacyl-tRNA that has been traditionally used as an inhibitor of protein synthesis since it specifically incorporates at the C-terminus of nascent polypeptide chains thereby stopping translation. Based on this observation, a new and significantly faster protein labeling approach has been developed:
C-terminal labeled full-length protein can be produced by in vitro translation in the presence of low concentrations of labeled dC-puromycin conjugates[1].
These proteins have been successfully used to analyze protein-protein interactions (pull-down assay, FCCS, protein-protein microarrays) and protein-DNA interactions (DNA microarrays)[2,3,4].
In contrast to traditionally applied posttranslational labeling approaches, additional time consuming purification steps can be avoided due to the synchronization of protein expression and labeling.
[1] Miyamoto-Sato et al. (2000) Specific bonding of puromycin to full-length protein at the C-terminus. Nucleic Acids Res. 28 (5):1176.
[2] Nemoto et al. (1999) Fluorescence labeling of the C-terminus of proteins with a puromycin analogue in cell-free translation systems. FEBS Letters 462:43.
[3] Doi et al. (2002) Novel Fluorescence Labeling and High-Throughput Assay Technologies for In Vitro Analysis of Protein Interactions. Genome Research 12:487.
[4] Kawahashi et al. (2007) High-throughput fluorescence labeling of full-length cDNA products based on a reconstituted translation system. J. Biochem. 141 (1):19.