cGAMPs are cyclic dinucleotides (CDNs) consisting of a 12-or-13-membered ring scaffold and two ribonucleotides. To date, two cGAMPs have been discovered in nature - 2‘,3‘-cGAMP[1] and 3‘,3‘-cGAMP[2].
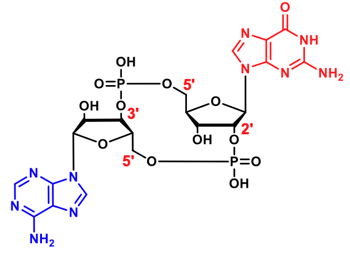
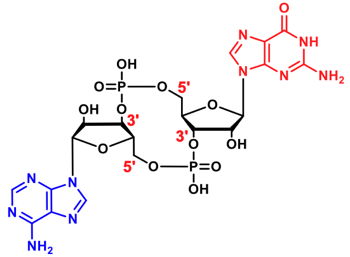
| Compound | Cat. No. | Amount | Price |
|---|---|---|---|
| 2',3'-cGAMP | NU-249S | 1 µmol | 96,43 € |
| NU-249L | 5 x 1 µmol | 385,70 € | |
| 3',3'-cGAMP | NU-986S | 1 µmol | 96,43 € |
| NU-986L | 5 x 1 µmol | 385,70 € |

2‘,3‘-cGAMP and 3‘,3‘-cGAMP in stock at Jena Bioscience. Please email Thomas for other cyclic dinucleotide pairs or with technical questions!
[1] Ablasser et al. (2013) cGAS produces a 2‘-5‘-linked cyclic dinucleotide second messenger that activates STING. Nature 498:380.
[2] Davies et al. (2012) Coordinated regulation of accessory genetic elements produces cyclic dinucleotides for V. cholerae virulence. Cell 149:358.
[3] Gao et al. (2013) Cyclic [G(2',5')pA(3',5')p] is the metazoan second messenger produced by DNA-activated cyclic GMP-AMP synthase. Cell 153 (5):1094.
[4] Tao et al. (2016) cGAS-cGAMP-STING: The three musketeers of cytosolic DNA sensing and signaling. IUBMB Life 68 (11):858.
[5] Opoku-Temeng et al. (2016) Cyclic dinucleotide (c-di-GMP, c-di-AMP, and cGAMP) signalings have come of age to be inhibited by small molecules. Chem. Commun. 52:9327.
[6] Ren et al. (2015) Structural basis for molecular discrimination by a 3‘,3‘-cGAMP sensing riboswitch. Cell Reports 11:1.