5’-capping and epigenetic nucleotide modifications are critical for efficient translation and reduced immunogenicity of synthetic mRNA[1-16].
The classic combination of modifications consists of ARCA, 5-methylcytidine and pseudouridine that are conveniently introduced by correspondingly modified nucleotides via T7 RNA polymerase-mediated in vitro transcription (Tab. 1, Fig. 1).
Promising extensions of the mRNA modification toolbox are available (e.g. N1-Methylpseudouridine[10], N4-Acetyl-Cytidine[5,6]) or the introduction of a Cap 1 moiety[15,16]). The optimal combination of Cap moiety & nucleotide modification however, needs to be individually determined for each mRNA target. Investigations can conveniently be performed with HighYield T7 mRNA Testkits (Tab. 1).
| Product | Application | Modified nucleotides/Cap analogs included | Alternative to |
|---|---|---|---|
| HighYield T7 mRNA Modification Testkit | Synthesis of differentially uridine-, cytidine- or adenosine-modified (m)RNA with Cap 0 or Cap 1 moiety | ARCA Cap 1 AG (3'-OMe) Pseudo-UTP, N1-Methylpseudo-UTP, 5-Methoxy-UTP, 2-Thio-UTP, 5-Methyl-CTP, N4-Acetyl-CTP, N6-Methyl-ATP, N1-Methyl-ATP |
n/a |
| HighYield T7 Uridine Modification Testkit | Synthesis of differentially uridine-modified (m)RNA with Cap 0 or Cap 1 moiety | ARCA Cap 1 AG (3'-OMe) Pseudo-UTP N1-Methylpseudo-UTP 5-Methoxy-UTP 2-Thio-UTP |
n/a |
| HighYield T7 Cytidine Modification Testkit | Synthesis of differentially cytidine-modified (m)RNA with Cap 0 or Cap 1 moiety | ARCA Cap 1 AG (3'-OMe) 5-Methyl-CTP N4-Acetyl-CTP |
n/a |
| HighYield T7 Cap Analog Testkit | Synthesis of Cap 0- and Cap 1-modified (m)RNA | ARCA Cap 1 AG (3'-OMe) |
n/a |
| HighYield T7 ARCA mRNA Synthesis Kit (m5CTP/Ψ-UTP) | Synthesis of Cap 0-, 5-methylcytosine & pseudouridine-modified mRNA | ARCA 5-Methyl-CTP Pseudo-UTP |
n/a |
| HighYield T7 ARCA mRNA Synthesis Kit | Synthesis of Cap 0-modified RNA | ARCA | HiScribe™ T7 ARCA mRNA Kit mMESSAGE mMACHINE™ T7 ULTRA Transcription Kit |
Check out our complete portfolio of HighYield T7 mRNA Synthesis Kits as well!
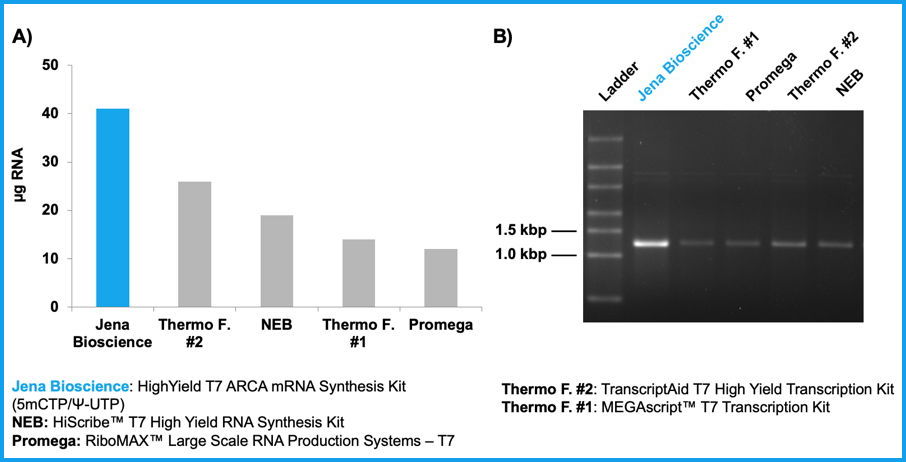
Figure 1: High yields of ARCA-, 5-methylcytidine- & pseudouridine-modified RNA are synthesized within 30 min using HighYield T7 ARCA mRNA Synthesis Kit (5mCTP/Ψ-UTP). 1.4 kbp RNA was generated using Jena Bioscience’s or competitor's in vitro transcription kits supplemented with ARCA, 5-Methyl-CTP and Pseudo-UTP (1 µg DNA template, 20 µl reaction, 30 min at 37 °C). Total NTP concentrations were used according to manufacturer’s instructions. Jena Bioscience/Thermo F. #2/Promega: 30 mM NTPs (6 mM ARCA, 1.5 mM GTP, 7.5 mM each ATP/UTP/CTP); Thermo F. #2/NEB: 40 mM NTPs (8 mM ARCA, 2 mM GTP, 10 mM each ATP/UTP/CTP).
A) RNA yield was determined by fluorescence measurement (100-fold dilution of transcription reactions, Quant-it™ RNA Assay Kit, broad range (Thermo F.)).
B) RNA integrity was analyzed by agarose gel electrophoresis (20-fold dilution of transcription reactions, equal volumes of each probe were run on a 1 % agarose/TAE gel supplemented with EtBr).

Please contact Barbara with all questions or inquiries you may have!
[1] Karikó et al. (2005) Suppression of RNA Recognition by Toll-like Receptors: The Impact of Nucleoside Modification and the Evolutionary Origin of RNA. Immunity 23:165.
[2] Karikó et al. (2008) Incorporation of Pseudouridine into mRNA Yields Superior Nonimmunogenic Vector With Increased Translational Capacity and Biological Stability. Mol. Ther. 16 (11):1833.
[3] Kormann et al. (2011) Expression of therapeutic proteins after delivery of chemically modified mRNA in mice. Nature Biotechnology 29 (2):154.
[4] Warren et al. (2011) Highly Efficient Reprogramming to Pluripotency and Directed Differentiation of Human Cells with Synthetic Modified mRNA. Cell Stem Cell 7:618.
[5] Svitkin et al. (2017) N1-methyl-pseudouridine in mRNA enhances translation through eIF2alpha-dependent and independent mechanisms by increasing ribosome density. Nucleic Acid Res 45 (10):6023.
[6] Andies et al. (2015) N1-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice. J. Control. Release 217:337.
[7] Li et al. (2016) Effects of Chemically Modified Messenger RNA on Protein Expression. Bioconjugate Chem. 27:849.
[8] Arango et al. (2018) Acetylation of Cytidine in mRNA Promotes Translation Efficiency. Cell 175 (7):1872.
[9] Sinclair et al. (2017) Profiling Cytidine Acetylation with Specific Affinity and Reactivity. ACS Chem. Neurosci. 12 (12):2922.
[10] Dominissini et al. (2016) The dynamic N1-methyladenosine methylome in eukaryotic messenger RNA. Nature 530:441.
[11] Wienert et al. (2018) In vitro transcribed guide RNAs trigger an innate immune response via RIG-I pathway. PLoS Biol. 16 (7):e2005840.
[12] Kim et al. (2018) CRISPR RNAs trigger innate immune responses in human cells. Genome Res. 28 (3):367.
[13] Badieyan et al. (2019) Concise Review: Application of Chemically Modified mRNA in Cell Fate Conversion and Tissue Engineering. Stem Cells Translational Medicine 8:833.
[14] Hadas et al. (2019) Optimizing Modified mRNA In Vitro Synthesis Protocol for Heart Gene Therapy. Molecular Therapy: Methods & Clinical Development 14:300.
[15] Shatkinet al. (1976) Capping of eukaryotic mRNAs. Cell 9 (4):645.
[16] Gallowayet al.(2019) mRNA cap regulation in mammalian cell function and fate. Biochimica et Biophysica Acta 1862 (3):270.