Most RNAs (both coding and non-coding) exert their functions in the form of complexes with RNA-binding proteins (RBPs)[1]. These complexes play crucial roles in post-transcriptional gene regulation processes, such as splicing, cleavage, polyadenylation, stabilization and translation of RNA[2,3].
The most successful methods so far for discovering RBP complexes are based on RNA-protein crosslinking using UV light irradiation (Photocrosslinking) and isolation of the resulting complexes by immunoprecipitation (CLIP)[8,9]. UV-irradiation lead to formation of stabile covalent bindings in between of RNAs and RNA-binding proteins affording the isolation of the complexes[5]. Photoactivatable ribonucleoside-Enhanced CLIP (PAR-CLIP) is an attractive further development involving incorporation of photoreactive nucleosides, such as 4-Thio-uridine (Figure 1A). It can be used with various cell types[1,6]. Moreover, the corresponding triphosphate 4-Thio-UTP is applicable for in vitro experiments identifying RBPs[4,7] (Figure 1B).
Due to equal base-pairing properties of 4-Thio-U and uridine, maximal RNA integrity is given. Since 4-Thio-U is activated at longer wavelenght than U (365 instead of 254 nm), less RNA damaging occurs[4,5].
There are many other benefits to using 4-Thio-uridine for discovering Protein-RNA interactions in cells or 4-Thio-UTP during your individual in vitro experiment:
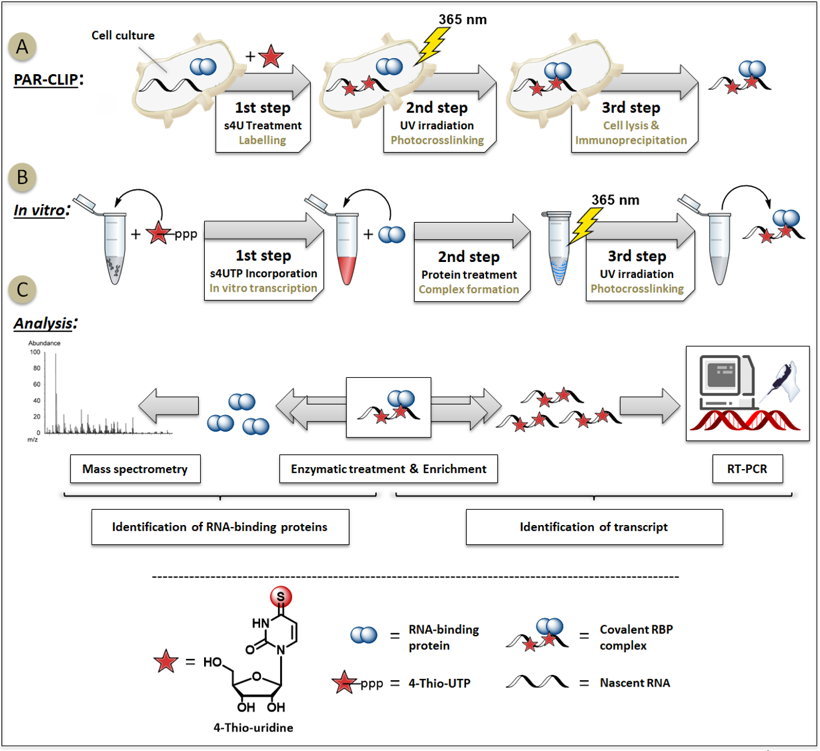
Figure 1: Schematic workflow of discovering RNA-protein interactions.
A) PAR-CLIP: Photoreactive 4-Thio-uridine is incorporated into nascent transcripts in cells, subsequent photocrosslinking of RNA to proteins (using 365 nm UV-light) prior to digestion[2-4].
B) Isolation of RBP complexes after in vitro transcription of 4-Thio-UTP and protein treatment[3,4,7].
C) Several enrichment and analysis strategies are possible to identify either RNA-binding proteins and its binding sites or RNA sequences[1-7].
Our HighYield T7 RNA Crosslinking Kit is containing T7 Polymerase that is validated for in vitro transcription of 4-Thio-UTP[7].
| Product | Cat.-No. | Amount | Price |
|---|---|---|---|
| 4-Thio-uridine | N-RP-2304-100 | 100 mg | 173,64 € |
| N-RP-2304-250 | 250 mg | 315,70 € | |
| 4-Thio-uridine-5'-triphosphate, Sodium salt (4-Thio-UTP) |
NU-1156S | 10 μl (100 mM) | 80,68 € |
| NU-1156L | 5 x 10 μl (100 mM) | 352,16 € | |
| HighYield T7 RNA Crosslinking Kit (4-Thio-UTP) | RNT-135 | 15 reactions x 20 µl | 295,00 € |

Please contact Barbara with all questions or inquiries you may have!
[1] Huang et al. (2018) Transcriptome-wide discovery of coding and noncoding RNA-binding proteins. PNAS 115 (17):E3879.
[2] Licatalosi et al. (2020) Approaches for measuring the dynamics of RNA–protein interactions. WIREs RNA 11:e1565.
[3] Ascano et al. (2012) Identification of RNA–protein interaction networks using PAR-CLIP. WIREs RNA 3:159.
[4] Sontheimer (1994) Site-specific RNA crosslinking with 4-thiouridine. Mol. Biol. Rep. 20 (1):35.
[5] He et al. (2016) High-Resolution Mapping of RNA-Binding Regions in the Nuclear Proteome of Embryonic Stem Cells. Mol. Cell 64 (2):416.
[6] Duffy et al. (2019) Gaining insight into transcriptome-wide RNA population dynamics through the chemistry of 4-thiouridine. WIREs RNA 10:e1513.
[7] Choi et al. (2017) An in vitro technique to identify the RNA binding-site sequences for RNA-binding proteins. BioTechniques 63 (1):28.
[8] Favre et al. (1986) 4-Thiouridine photosensitized RNA-protein crosslinking in mammalian cells. Biochem. Biophys. Res. Commun. 141 (2):847.
[9] Hajnsdorf et al. (1987) RNA protein crosslinks introduced into E. coli ribosomes by use of the intrinsic probe 4-thiouridine. Photochem. Photobiol. 45 (4):445.